By Richard Morimoto
The accumulation of misfolded protein marks the accrual of years as the body ages. Could heat shock proteins be used to reduce the effects of aging and diminish the risk of disease by untangling improperly folded proteins?
What does a molecular thermometer look like? This seemed to be a simple question, not much different from the many science fair projects I had done in grade school and high school in Chicago. But rather than the simple solutions I’d present on triptych posterboards, the answer to this question has kept me fascinated for my entire career. The cell’s thermometer appears to be a network of stress-sensing transcription factors and specialized proteins—molecular chaperones—that function as the guardian of the proteome, sensing damage and keeping the cell’s proteins properly folded as they roll off the production line of ribosomes. Exciting as that was, none of us working on this question at the time could have predicted that this thermometer might also control the body’s fountain of youth and provide new ways of thinking about disease.
Proteins are fundamental building blocks for the cell; they are the predominant products of the genome that provide much of the shape and functionality of cells, tissues, and organisms. The proper synthesis, folding, assembly, translocation, and clearance of proteins is essential for the health of the cell and the organism. Proteins also provide the essential parts to replenish molecular machines for biosynthetic processes and ensure their efficient functioning in the adult cell, a process critical for longevity. At the root of the problem is a fundamental process: protein folding. When quality control—as overseen by heat shock proteins and molecular chaperones—slips, errors occur and persist. This interferes with molecular processes, which can lead to disease. When these events occur in neurons, the consequences can be devastating, leading to major classes of neurological disorders, like multiple sclerosis, Huntington’s disease, Parkinson’s disease, and Alzheimer’s disease.
 I first became intrigued by heat shock proteins while listening to a seminar by Matt Meselson, who was visiting the University of Chicago to receive an honorary doctorate in the late 1970s. At the time, I had just begun my second year as a graduate student in Murray Rabinowitz’s laboratory in the Department of Biochemistry. While the seminar piqued my interest in this newly discovered heat shock response, it was through my subsequent conversations with Susan Lindquist—who had completed her research with Meselson and had just recently arrived on campus as a postdoctoral fellow—that I really began to appreciate the complexities of this biological response to stress.
I first became intrigued by heat shock proteins while listening to a seminar by Matt Meselson, who was visiting the University of Chicago to receive an honorary doctorate in the late 1970s. At the time, I had just begun my second year as a graduate student in Murray Rabinowitz’s laboratory in the Department of Biochemistry. While the seminar piqued my interest in this newly discovered heat shock response, it was through my subsequent conversations with Susan Lindquist—who had completed her research with Meselson and had just recently arrived on campus as a postdoctoral fellow—that I really began to appreciate the complexities of this biological response to stress.
In between my efforts to complete my thesis research on the yeast mitochondrial genome and compete in intramural sports with our team, the Kimwipes, I made time to learn more from Lindquist about the heat shock response. Discovered in 1962 by Ferruccio Ritossa, the heat shock response was described as the temperature-induced change in the organization of the tightly packed Drosophila salivary gland chromosome, leading to the appearance of puffs.
Ritossa’s “heat shock” puffs, which by light microscopy looked like cotton balls compressed between sections of tightly packed chromosomes, were also induced by exposure to dinitrophenol, ethanol, and salicylate. However, it was the demonstration that these puffs were new sites of transcription that was most impressive—Ritossa could detect newly synthesized RNA within minutes of puffing. The significance of this was soon revealed by Lindquist and Steven Henikoff in the Meselson lab and Allan Spradling with Mary-Lou Pardue at MIT. These heat shock–induced messenger RNAs encoded a set of proteins, now widely known as the heat shock proteins. These are widely studied as Hsp90, Hsp70, and a family of proteins that guide conformation and folding, and prevent misfolding in cells of all species.
Using a clever trick of cellular biochemistry to enrich for heat shock messenger RNA, the laboratories of David Hogness at Stanford, Walter Gehring at University of Basel, Brian McCarthy at UCSF, Alfred Tissieres at the University of Geneva, and Meselson all cloned the first heat shock genes from Drosophila in the late 1970s and early 1980s. 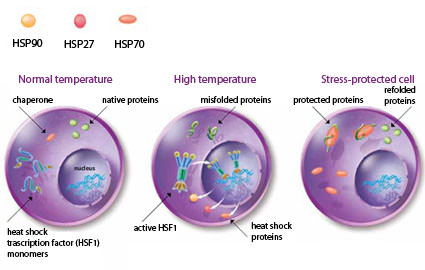
Heat shock: Exposure of cells to heat shock or other forms of physiological stress elevates the level of misfolded proteins. Accumulation of damaged proteins is prevented by induction of the heat shock response and the expression of heat shock proteins (Hsp90, Hsp70, and Hsp27). Cells at normal temperature have mostly native proteins and heat shock transcription factors in an inactive state. Upon heat shock, proteins misfold and heat shock factors are activated and result in the elevated expression of heat shock genes. In the stress-protected cell, heat shock proteins stabilize misfolded proteins, prevent the formation of aggregates, and enhance the levels of native functional proteins.
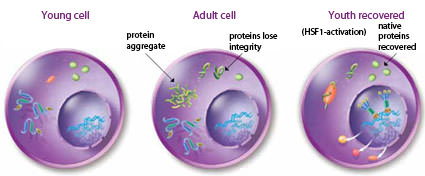
Heat shock factors in aging: As the cells in an organism age, proteins lose their integrity and appear as misfolded molecules that can aggregate. In the young cell, proteins are mostly in the native folded state. During aging, misfolded proteins appear and can form protein aggregates that interfere with cellular function. The youth recovered state can be attained by enhanced activity of heat shock transcription factor to elevate the levels of heat shock proteins in the cell, thus reducing the appearance of damaged proteins.
By this time, I had received my PhD from Chicago and was a postdoc with Matt Meselson in the Harvard biolabs. During this period, I became intrigued with the possibility that heat shock genes might be expressed in other eukaryotes. The counter opinion at the time was that heat shock response was a fruit fly thing. While in hindsight this might seem an odd and naïve question, our knowledge in the early 1980s was very limited and there was no reason to assume that a temperature response in fruit flies should have anything to do with humans. Moreover, the concepts of heat shock proteins as molecular chaperones would not appear for nearly another decade. Nevertheless, armed with the newly developed method of Southern blotting, I decided to search for the heat shock genes in other eukaryotes. Being located in the Harvard biolabs where there was an impressive collection of organisms under investigation, it was just one day’s work to wander the long halls with an ice bucket and return with genomic DNA from a wide array of life’s creatures. After running my collection of DNA on an agarose gel and incubating the nitrocellulose transfer paper with my radiolabeled Hsp70 gene fragments, I searched for the DNA fragments that were complementary.
I still remember the water dripping down my arm as I pulled out the developed X-ray film and saw bands of different sizes in each of the lanes of the gel. Hsp70 was everywhere and in all eukaryotic genomes, including humans’. The characterization of the human Hsp70 gene then became the focus of my new laboratory in the Department of Biochemistry, Molecular Biology, and Cell Biology at Northwestern University in Evanston, Ill. Around the same time, Jim Bardwell and Betty Craig at the University of Wisconsin at Madison had identified a gene in E. coli, dnaK, which was the bacterial equivalent of Hsp70, leading to the realization that heat shock genes were essentially identical from bacteria to humans and subsequently shown to be present in all five kingdoms.1
None of us working on this question at the time could have predicted that this thermometer might also control the body’s fountain of youth and provide new ways of thinking about disease.
With the cloning of human Hsp70 and subsequently other members of the human Hsp70 gene family, our efforts at Northwestern initially focused on learning how this gene was regulated in fruit flies. While looking for sequence homologies across species, we spent a lot of time looking at the regulatory regions of the human heat shock genes that would be bound by the heat shock transcription factor. Efforts mostly from Carl Wu at NIH, Bob Kingston at Harvard, John Lis at Cornell, and my lab led to the identification of a family of heat shock transcription factors that controlled the first step of the heat shock response. Was this the molecular thermometer that I had been searching for? These heat shock factors changed their shape when the temperature changed, binding to the heat shock gene promoters and initiating the expression of heat shock genes (see graphic below). Moreover, humans expressed three such heat shock factor genes—as opposed to one in yeast, Drosophila, and C. elegans—suggesting that these genes, with such exquisite regulation, might have other functions that we had yet to discover.
 Additional insights on the molecular thermometer came from studies showing that the heat shock proteins were regulating the very transcription factor that initiated their production in the nucleus. The function of heat shock proteins as molecular chaperones to guide folding, to ensure alternate conformational states, and to recognize and suppress misfolding was becoming well established. Therefore, the cellular thermometer wasn’t just measuring temperature, it also monitored for the appearance of damaged proteins. Heat shock and other stressors would lead to a flux of misfolded and damaged proteins, shifting the chaperone equilibrium in the cell toward capturing these damaged proteins and releasing Hsf1 to self-associate and form trimers capable of binding to the heat shock elements in the genome. The chaperones, therefore, had yet another role: to repress the transcriptional activity of Hsf1, providing a feedback loop for the heat shock response when sufficient levels of heat shock proteins were available.
Additional insights on the molecular thermometer came from studies showing that the heat shock proteins were regulating the very transcription factor that initiated their production in the nucleus. The function of heat shock proteins as molecular chaperones to guide folding, to ensure alternate conformational states, and to recognize and suppress misfolding was becoming well established. Therefore, the cellular thermometer wasn’t just measuring temperature, it also monitored for the appearance of damaged proteins. Heat shock and other stressors would lead to a flux of misfolded and damaged proteins, shifting the chaperone equilibrium in the cell toward capturing these damaged proteins and releasing Hsf1 to self-associate and form trimers capable of binding to the heat shock elements in the genome. The chaperones, therefore, had yet another role: to repress the transcriptional activity of Hsf1, providing a feedback loop for the heat shock response when sufficient levels of heat shock proteins were available.
As my lab started exploring the normal functions of heat shock proteins in the cell, I began to feel somewhat constrained by the human tissue culture cell lines we had been using. While the cells allowed us to probe the identity and function of the relevant molecules, it was unclear how these changes related to a whole organism. In around 1999, we introduced C. elegans into our lab as a new model system to address a question on the regulation of the heat shock response. We drew inspiration from a 1994 paper by Max Perutz showing that a mutation in polyglutamine that caused the protein to aggregate was responsible for Huntington’s disease, a profound and devastating neurodegenerative disease. In short time, C. elegans became our system of choice for exploring new stress signals such as the expression of expanded polyglutamine proteins.2 Could misfolded proteins, as prompted by the expression of polyglutamine, provide insights into the molecular thermometer?
Buoyed by these possibilities, I submitted the renewal for my long-standing NIH grant and was nominated for a Merit Award. This gift of 10-year support from the National Institute of General Medical Sciences (NIGMS) came at exactly the right time. In addition to developing new animal models for expression of polyglutamine expansion in neurons and muscle cells, we observed that the aggregation and associated toxicity of these proteins was proportional to their length and also to the age of the animal. This led us to ask whether suppressing the insulin signalling pathways that regulate lifespan might affect the aggregation of polyglutamine. The results showed a clear relationship,3 moreover subsequent studies revealed that Hsf1 was essential for lifespan via the insulin signalling pathway. These observations revealed that the heat shock response was not only important for survival of acute stress but equally for day-to-day events of protein toxicity that could affect the health of the cell’s proteins and lifespan.
By this point, an increasing amount of work in our lab had shifted toward the use of C. elegans, and this whole organismal thinking gave me leeway to probe beyond the concept of stress at a cellular level to begin to address how the animal senses and integrates external information at the molecular level. If stress affected aging, then perhaps these heat shock proteins that sensed and protected the cell from accumulated damage might help slow the effects of aging.
Ritossa’s “heat shock” puffs, which by light microscopy looked like cotton balls compressed between sections of tightly packed chromosomes, proved to be new sites of transcription.
Because we had earlier observed that many diseases associated with aggregation-prone proteins exhibit age-related phenotypes and that all protein misfolding diseases are associated with aging, it seemed logical to ask whether temperature-sensitive metastable proteins lost function during aging. We had observed earlier that polyglutamine expressed in different tissues of C. elegans caused temperature-sensitive proteins in the same tissue to completely misfold and aggregate. This result really turned our heads, for none of us had expected that a single aggregation-prone protein would have such global effects! Indeed, when the same temperature-sensitive proteins were followed during aging alone, we observed that they all misfolded and lost function early on in natural aging.4 When I saw these results, I was shocked. It suggested that the machines that keep proteins folded properly would have already started to fall apart as an organism reaches maturity. Returning to Hsf1, we learned that the heat shock response and other cytoprotective responses were compromised during aging. However, despite the rather profound consequences of these observations, we found that activation of Hsf1 in early development could suppress this collapse of protein-folding homeostasis (proteostasis). The implications are intriguing. While it is always possible that such observations do not extend to more complex metazoans, we are all made of proteins and it seems likely that the tales of worms may help us understand human aging.
But there was more to the story. Our recent work suggests there may be ways of modulating proteostasis that don’t involve interfering with Hsf1 directly. We decided to plunge into a classical genetic screen of C. elegans mutants that demonstrated protein misfolding in the muscle cells. While one might have expected to get many of the same genes already identified, we found something completely new. The mutation that affected misfolding in the muscle cell corresponded to a transcription factor that controls the production of the neurotransmitter GABA, responsible for mediating excitatory neurotransmitters and controlling muscle tone! Remarkably, any imbalance in the GABA pathway leading to overstimulation caused the postsynaptic muscle cell to think it was stressed and to start misfolding proteins.5 Suddenly, the molecular thermometer got more complicated with the realization that the activity of neurons could affect the thermometer of another cell.
Many human diseases thought to be unrelated may share commonalities of defects in protein folding homeostasis.
If this mechanism also occurs in humans, we may be able to find the neuronal signature that controls protein misfolding in cells and activate the heat shock proteins in the skeletal muscle system to help restore function in muscular dystrophy and other motor neuron diseases. The findings also tease the possibility that there might be a way to control, stall, or regulate the misfolding associated with aging via a neurological mechanism. However, the story with neurons seems to be more nuanced. We observed that two neurons in C. elegans control the regulation of the heat shock response in all adult somatic cells. The role of active neuronal signalling and feedback control would seem to provide a basis by which cells and tissues activate a heat shock response according to need. This makes sense, as different tissues will have varied protein biosynthetic needs and environmental exposures. A “one-size-fits-all” approach to the heat shock response would not work.
With these exciting ideas that our research has led us to ponder, I and several of my colleagues (Jeff Kelly at The Scripps Research Institute and Andy Dillin at the Salk Institute) cofounded Proteostasis Therapeutics in Cambridge, Mass., to discover small molecule therapeutics that could correct the effects of misfolded proteins in disease. The approach is not without risk—the level of heat shock proteins are increased in many cancers—but the promise of the technology is certainly too exciting not to explore.
The heat shock response is a good story. It has humble beginnings of pure curiosity. Starting with a small band of “heat shockers,” the field has grown immensely and encompasses a multitude of disciplines and approaches fundamental to biology. In it are all of the elements of intrigue and surprise, with a remarkable cast of characters. Who would have predicted the chance observation of Ritossa’s that chromosomal puffs in Drosophila, induced by elevated temperature, would lead to discoveries across all of biology? Well beyond the satisfaction of these observations alone has been the recognition that many human diseases thought to be unrelated may share commonalities of defects in protein folding homeostasis and that the correction of these defects could have broad-reaching global effects on proteome stability and the health of the cell. All throughout this, I have been fortunate to have a wonderful family—my wife Joyce and children Emiko and Kenji and a fantastic set of colleagues and lab members. Perhaps the efforts to apply the discoveries of heat shock proteins successfully to maintaining health and curing disease is one way to thank them for their enthusiastic support.
BY RICHARD MORIMOTO
Richard Morimoto is the Bill and Gayle Cook Professor of Biology and director of the Rice Institute for Biomedical Research at Northwestern University, and a cofounder of the Proteostasis Therapeutics, Inc. in Cambridge, Mass.
References:
1. C. Hunt, R.I. Morimoto, “Conserved features of eukaryotic hsp70 genes revealed by comparison with the nucleotide sequence of human hsp70,” Proc Natl Acad Sci USA, 82:6455–59, 1985.
2. S.H. Satyal et al., “Polyglutamine aggregates alter protein folding homeostasis in Caenorhabditis elegans,” Proc Natl Acad Sci USA, 97:5750–55, 2000.
3. J.F. Morley et al., “The threshold for polyglutamine-expansion protein aggregation and cellular toxicity is dynamic and influenced by aging in Caenorhabditis elegans,” Proc Natl Acad Sci USA, 99:10417–22, 2002.
4. A. Ben-Zvi et al., “Collapse of proteostasis represents an early molecular event in Caenorhabditis elegans aging,” Proc Natl Acad Sci USA, 106:14914–19, 2009.
5. S. Garcia et al., “Neuronal signaling modulates protein homeostasis in Caenorhabditis elegans postsynaptic muscle cells,” Genes Dev, 21:3006–16, 2007. PMCID: PMC2049200
Read more: Shock and Age – The Scientist – Magazine of the Life Sciences http://www.the-scientist.com/article/display/57461/#ixzz0qs5xHTP4