A new technique developed at Rensselaer Polytechnic Institute allows researchers to collect large amounts of biochemical information from nanoscale bone samples. Along with adding important new insights into the fight against osteoporosis, this innovation opens up an entirely new proteomics-based approach to analyzing bone quality. It could even aid the archeological and forensic study of human skeletons.
“We’re able to take very small, nanoscale-sized bone samples, and determine the protein signatures of the bone,” said Deepak Vashishth, head of the Department of Biomedical Engineering at Rensselaer, who led the study. “This is a relatively quick, easy way for us to determine the history of the bone — how and when it formed — as well as the quality of the bone, and its likelihood to fracture.”
Results of the study, titled “Biochemical Characterization of Major Bone-Matrix Proteins Using Nanoscale-Size Bone Samples and Proteomics Methodology,” were released online in late May by the journal Molecular & Cellular Proteomics. The journal, published by the American Society for Biochemistry and Molecular Biology, will also feature the paper in an upcoming print edition.
The research, funded by the U.S. National Institutes of Health, was conducted in the laboratories of the Center for Biotechnology and Interdisciplinary Studies at Rensselaer.
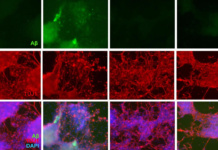
Bones are primarily composed of mineral, with the remaining amount composed of organic material. The vast majority of the organic material is collagen. The remaining non-collagenous organic material is a mixture of other proteins, which form an interlinked matrix. The quality of this matrix varies greatly with age, nutrition, and disease. Vashishth and his research group investigate this bone matrix to determine how the interaction and modification of individual proteins impact the development, structure, and strength of the overall bone.
In this study, they paired laser-capture microscopy with several other techniques to create an entirely new method for analyzing bone matrix. The analysis yields data about the concentration of different proteins in the bone matrix, which in turn leads to key information about the bone — such as when it was formed, how it has been modified, and if it is more or less prone to fracture.
Vashishth said this is an important step toward augmenting current osteoporosis diagnosis techniques, which measure bone loss and the quantity of bone present, with new, minimally invasive, proteomics-driven techniques for assessing the quality of the bone.
The young field of proteomics focuses on the structure and function of proteins, and is ripe for innovation, Vashishth said. The term “proteomics” echoes the word genomics, the study of genes. Proteomics seeks to decode the human proteome by documenting the structure, function, and interactions of proteins.
“This is kind of a new area, because bone fracture has always been looked at from a bone calcium perspective, a mineral perspective, and current osteoporosis treatment methods are all geared toward that,” he said. “In osteoporosis, very little attention has been paid to bone proteins. That’s why we’re very excited about our new proteomics-based method to read a bone’s protein signature, and assess the quality of the bone. I think it opens up a new avenue for approaching and studying osteoporosis.”
Like all tissues in the human body, bones regenerate themselves over time. Bones regenerate much slower than other tissues, however, and the skeleton takes about 10 years to gradually replace itself with new tissue. Different parts of a bone regenerate at different rates, meaning some areas of a bone may be older and more susceptible to fracture, while other areas of the same bone are newer and sturdier. Older and younger parts of a bone have different protein signatures and react differently to medical treatments. Vashishth said his new method is an easy way to help differentiate between different aged areas of bone, determine their quality, and forecast their susceptibility to fracture.
Finally, along with pushing forward the emerging field of bone proteomics and opening up new possibilities for studying and treating osteoporosis, Vashishth’s findings could prove useful to researchers in other areas who deal with bone. Forensics, biology, anthropology, archaeology, and other areas where bone samples are truly rare, small, and precious would likely find it useful to analyze bone protein signatures with minimal damage to the bone sample, he said. This protein signature information could offer new insight into how bones were formed, along with the nutrition and diet of those individuals.
Co-authors of the study are Wilfredo Colon, professor in the Rensselaer Department of Chemistry and Biological Chemistry; as well as postdoctoral researcher Grazyna Sroga and doctoral student Lamya Karim, both in the Rensselaer Department of Biomedical Engineering.
Story Source:
The above story is reprinted (with editorial adaptations by ScienceDaily staff) from materials provided by Rensselaer Polytechnic Institute.
Journal Reference:
- G. E. Sroga, L. Karim, W. Colon, D. Vashishth. Biochemical characterization of major bone-matrix proteins using nanoscale-size bone samples and proteomics methodology. Molecular & Cellular Proteomics, 2011; DOI: 10.1074/mcp.M110.006718